GE2R
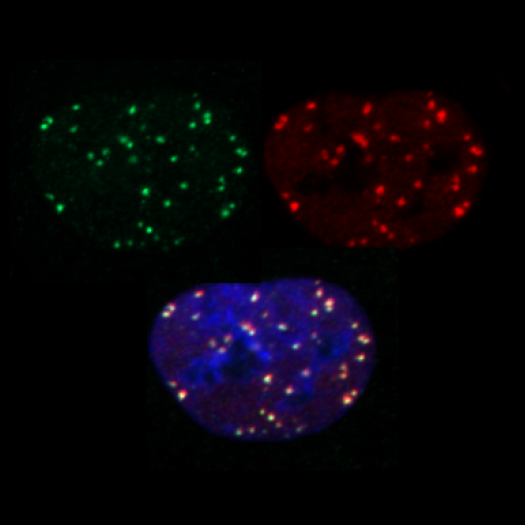
marquage yH2AX, 53BP1 et merge en tryptique


marquage yH2AX, 53BP1 et merge en tryptique
Edition du génome, réparation de l’ADN et réponses cellulaires
Les cellules eucaryotes ont développé diverses voies de réparation pour résoudre les dommages sur l’ADN et la réparation fidèle est cruciale pour maintenir l’intégrité du génome. D’autre part, la modification du génome avec des nucléases à façon tire parti de ces voies de réparation pour produire des modifications génomiques permanentes et est désormais une stratégie majeure pour étudier l’organisation du génome et la fonction des gènes, créer des modèles animaux de maladies et améliorer la thérapie génique.
Notre équipe s’intéresse à l’étude des voies de réparation de l’ADN et leurs rôles dans la modification du génome ainsi que dans différentes situations biologiques, en tirant parti des nouvelles possibilités offertes par les nucléases à façon.
Nos projets s’articulent autour de deux grands axes :
Les cellules ont la capacité de traiter un large éventail de lésions de l’ADN, et l’édition du génome, qui commence toujours par l’introduction d’une lésion ciblée, dépend ensuite de la réparation de l’ADN cellulaire pour parvenir à une modification du génome. L’objectif des approches d’édition du génome est de produire une séquence désirée à des taux élevés, avec très peu d’autres mutations.
Ces dernières années, les nucléases CRISPR-Cas ont été utilisées pour éditer le génome après l’introduction d’une cassure double brin (DSB) au site ciblé puis réparation, soit par réparation dirigée par homologies (HDR), soit par réparation par jonction d’extrémités. La modification précise du génome, c’est-à-dire la modification programmée de séquence, peut être réalisée après HDR, en copiant un ADN « modèle » portant le changement de séquence souhaité (appelé ADN donneur) et introduit en même temps que la nucléase. Cependant, les voies de réparation par jonction d’extrémités, la jonction d’extrémités non homologue canonique (cNHEJ) et la jonction d’extrémités par microhomologies (MMEJ), dominent généralement sur la voie HDR, rendant la modification du génome par la voie HDR peu efficace dans de nombreux systèmes. La manipulation des voies de réparation des DSB représente une des approches pour obtenir la séquence désirée à des taux importants par rapport à d’autres modifications.
Une deuxième génération d’approches d’édition du génome a été décrite ; il s’agit des approches d’édition de base et de « Prime editing », qui permettent une modification du génome sans cassure double brin et sans ADN donneur. Elles sont basées sur l’utilisation de protéines de fusion entre une cas9 nickase et une désaminase, ou une transcriptase inverse associée à un ARN guide modifié (appelé pegARN, pour ARN guide de Prime editing), respectivement. Elles permettent d’obtenir des changements de séquence efficaces et précis dans certaines conditions : pour les éditeurs de base, la précision dépend principalement de la séquence cible, tandis que pour les Prime éditeurs (PE), l’efficacité varie fortement selon les types cellulaires et les types de modifications.
Un des objectifs de notre équipe est d’augmenter l’efficacité de la modification précise du génome et de mieux comprendre et contrôler les mécanismes de réparation de l’ADN impliqués. Nous avons travaillé sur le ciblage au niveau de la cassure de protéines impliquées dans la voie HDR afin de favoriser cette voie (6). Nos objectifs actuels sont de : (i) caractériser de nouveaux acteurs de la voie MMEJ qui semblent jouer un rôle prédominant dans la réparation des DSB infligées par les nucléases CRISPR (11) et de les exploiter pour améliorer la modification du génome ; (ii) surmonter les limitations actuelles de l’approche de Prime Editing.
Ces dernières années, nous avons développé des outils d’édition du génome et les avons utilisés dans différents systèmes biologiques : dans différents modèles animaux pour des études fonctionnelles (2) et dans des cellules humaines pour la thérapie génique, et nous proposons ces outils aux laboratoires à travers notre plateforme TACGENE.

Figure 1: Stratégies d’édition du génome avec les systèmes CRISPR-Cas.
Les différentes approches de modification du génome, les voies de réparation de l'ADN impliquées et les types de modifications réalisées sont indiqués (MMR : Réparation des mésappariements, BER : Réparation par excision de base, HDR : Réparation dirigée par homologies ; MMEJ : Jonction d'extrémités par microhomologies ; NHEJ : Jonction d'extrémités non homologue).
Nous avons choisi d’étendre nos études sur la réparation de l’ADN aux tardigrades, un nouvel organisme modèle in vivo connu pour sa tolérance aux environnements extrêmes. Les tardigrades sont résistants à une grande variété d’environnements extrêmes, y compris des conditions qui induisent des niveaux élevés de dommages à l’ADN. L’étude des tardigrades pourrait donc permettre l’identification de nouvelles protéines et mécanismes pour gérer les dommages à l’ADN. De manière intéressante, une protéine de tardigrade appelée Dsup, qui interagit avec les nucléosomes et peut les protéger contre les dommages à l’ADN, a été identifiée et est probablement impliquée dans cette résistance extrême. Cependant, le gène Dsup n’est présent que chez certains tardigrades (dans la super-famille des Hypsibiidae), suggérant que d’autres gènes et mécanismes pourraient être impliqués.
Notre objectif est d’examiner les mécanismes moléculaires de résistance des tardigrades aux dommages de l’ADN, ainsi que la spécificité cellulaire de ces mécanismes, en identifiant les gènes candidats et en analysant la régulation des voies de réparation de l’ADN dans différentes espèces de tardigrades, par des approches de génomique fonctionnelle et comparative. Nous explorerons également les applications possibles des protéines de tardigrade dans des organismes hétérologues.
Par l’analyse du transcriptome en réponse aux radiations ionisantes dans trois espèces de tardigrades, nous avons récemment identifié un nouveau gène que nous avons nommé « gène de réponse aux dommages à l’ADN de tardigrade 1 » (TDR1) qui n’est présent que chez les tardigrades et pourrait jouer un rôle clé dans leur résistance aux dommages de l’ADN. Nous avons donc commencé à caractériser ses mécanismes d’action (1).
Enfin, plusieurs formes de reproduction ont été rapportées chez les tardigrades, la reproduction asexuée par parthénogenèse ou la reproduction sexuée, et nous proposons d’explorer cet aspect fascinant de la biologie des tardigrades et ses conséquences sur l’adaptation des tardigrades, en utilisant des approches de génomique comparative, associées à la caractérisation cellulaire et moléculaire du développement des cellules germinales et de la méiose.

Figure 2 : Localisation nucléaire de TDR1 chez les tardigrades Hypsibius exemplaris.
Les plasmides d'expression de la mCherry et de HeTDR1-mNeonGreen sous le contrôle du promoteur HeActin ont été introduits chez des tardigrades adultes H. exemplaris. L'imagerie confocale 8 jours post-introduction montre la localisation nucléaire de la protéine de fusion HeTDR1-mNeonGreen.
M. Anoud, E. Delagoutte, Q. Helleu, A. Brion, E. Duvernois-Berthet, M. As, X. Marques, K. Lamribet, C. Senamaud, L. Jourdren, A. Adrait, S. Heinrich, G. Toutirais, S. Hamlaoui, G. Gropplero, I. Giovannini, L. Ponger, M. Gèze, C. Blugeon, Y. Coute, R. Guidetti, L Rebecchi, C. Giovannangeli, A. De Cian, J-P. Concordet. Comparative transcriptomics reveal a novel tardigrade specific DNA binding protein induced in response to ionizing radiation, eLife (accepted), bioRxiv 2023.09.08.556854; doi: https://doi.org/10.1101/2023.09.08.556854
Rosello M, Serafini M, Mignani L, Finazzi D, Giovannangeli C, Mione MC, Concordet JP, Del Bene FC. Disease modeling by efficient genome editing using a near PAM-less base editor in vivo. Nat Commun. 2022 Jun 15;13(1):3435. doi: 10.1038/s41467-022-31172-z. PMID: 35701478
Geny S, Pichard S, Poterszman A, Concordet JP. Gene Tagging with the CRISPR-Cas9 System to Facilitate Macromolecular Complex Purification. Methods Mol Biol. 2021; 2305:153-174. doi: 10.1007/978-1-0716-1406-8_8. PMID: 33950389.
Weber L, Frati G, Felix T, Hardouin G, Casini A, Wollenschlaeger C, Meneghini V, Masson C, De Cian A, Chalumeau A, Mavilio F, Amendola M, Andre-Schmutz I, Cereseto A, El Nemer W, Concordet JP, Giovannangeli C, Cavazzana M, Miccio A. Editing a γ-globin repressor binding site restores fetal hemoglobin synthesis and corrects the sickle cell disease phenotype. Sci Adv. 2020 Feb 12;6(7):eaay9392. doi: 10.1126/sciadv.aay9392. PMID: 32917636; PMCID: PMC7015694.
Concordet JP, Haeussler M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 2018 Jul 2;46(W1):W242-W245. doi: 10.1093/nar/gky354. PMID: 29762716; PMCID: PMC6030908.
Charpentier M, Khedher AHY, Menoret S, Brion A, Lamribet K, Dardillac E, Boix C, Perrouault L, Tesson L, Geny S, De Cian A, Itier JM, Anegon I, Lopez B, Giovannangeli C, Concordet JP. CtIP fusion to Cas9 enhances transgene integration by homology-dependent repair. Nat Commun. 2018 Mar 19;9(1):1133. doi: 10.1038/s41467-018-03475-7. PMID: 29556040; PMCID: PMC5859065.
Hosseini ES, Nikkhah M, Hamidieh AA, Fearnhead HO, Concordet JPC, Hosseinkhani S. The Lumiptosome, an engineered luminescent form of the apoptosome can report cell death by using the same Apaf-1 dependent pathway. J Cell Sci. 2020 May 27;133(10):jcs242636. doi: 10.1242/jcs.242636. PMID: 32461338.
de Oliveira VC, Gomes Mariano Junior C, Belizário JE, Krieger JE, Fernandes Bressan F, Roballo KCS, Fantinato-Neto P, Meirelles FV, Chiaratti MR, Concordet JP, Ambrósio CE. Characterization of post-edited cells modified in the TFAM gene by CRISPR/Cas9 technology in the bovine model. PLoS One. 2020 Jul 10;15(7):e0235856. doi: 10.1371/journal.pone.0235856. PMID: 32649732; PMCID: PMC7351154.
Babin L, Piganeau M, Renouf B, Lamribet K, Thirant C, Deriano L, Mercher T, Giovannangeli C, Brunet EC. Chromosomal Translocation Formation Is Sufficient to Produce Fusion Circular RNAs Specific to Patient Tumor Cells. iScience. 2018 Jul 27;5:19-29. doi: 10.1016/j.isci.2018.06.007. Epub 2018 Jun 19. PMID: 30240643; PMCID: PMC6123901.
Concordet JP, Haeussler M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 2018 Jul 2;46(W1):W242-W245. doi: 10.1093/nar/gky354. PMID: 29762716; PMCID: PMC6030908.
Momose T, De Cian A, Shiba K, Inaba K, Giovannangeli C, Concordet JP. High doses of CRISPR/Cas9 ribonucleoprotein efficiently induce gene knockout with low mosaicism in the hydrozoan Clytia hemisphaerica through microhomology-mediated deletion. Sci Rep. 2018 Aug 6;8(1):11734. doi: 10.1038/s41598-018-30188-0. PMID: 30082705; PMCID: PMC6078951.
Renaud J-B., Boix C., Charpentier M., De Cian A., Cochennec J., Duvernois-Berthet E., Perrouault L., Tesson L., Edouard J., Thinard R., Cherifi Y., Menoret S., Fontanière S., de Crozé N., Fraichard A., Sohm F., Anegon I., Concordet J-P. & Giovannangeli C. (2016) Improved Genome Editing Efficiency and Flexibility Using Modified Oligonucleotides with TALEN and CRISPR-Cas9 Nucleases. Cell Reports. 14,9:2263-72.